Nonparametric ANOVA: Kruskal-Wallis Test.
Il a été proposé par Frank Wilcoxon en 1945 et par Henry Mann et Donald Ransom Whitney en 1947. Two random variables x and y are called independent if the probability distribution of one variable is not affected by the presence of another. It is considered to be the nonparametric equivalent to the two sample t-test. The Mann-Whitney U test and the Kruskal-Wallis test are nonparametric methods designed to detect whether 2 or more samples come from the same distribution or to test whether medians between comparison groups are different, under the assumption that the shapes of the underlying distributions are the same. Synonymous: Mann-Whitney test, Mann-Whitney U test, Wilcoxon-Mann-Whitney test and two-sample Wilcoxon test. With this same command, we can adjust the p-values according to a variety of methods. If you want continuity correction in wilcox.test just use argument correct=T.
The (Wilcoxon-) Mann-Whitney (WMW) test is the non-parametric equivalent of a pooled 2-Sample t-test. and added to the plots using the stat_compare_means() function. If not, the Mann-Whitney U two-sample test is not suitable since there is no specified direction of the hypothesized difference. Boxplots were generated using the geom box - plot() function. Groups with distribution deviating from normal were compared using the Mann–Whitney test (wilcox.test). data: The data to be displayed in this layer. You must supply mapping if there is no plot mapping. It provides an easier syntax to generate information-rich plots for statistical analysis of continuous (violin plots, scatterplots, histograms, dot plots, dot-and-whisker plots) or categorical (pie and bar charts) data. Some implementations use the exact distribution for small samples with the Wilcoxon-Mann-Whitney but not for the Kruskal-Wallis (yielding not-so-accurate p-values with small samples). For each candidate probe-gene pair, the Mann-Whitney U test is used to test the null hypothesis that overall gene expression in group M is greater than or equal than that in group U. It's also possible to perform the test for multiple response variables at the same time.
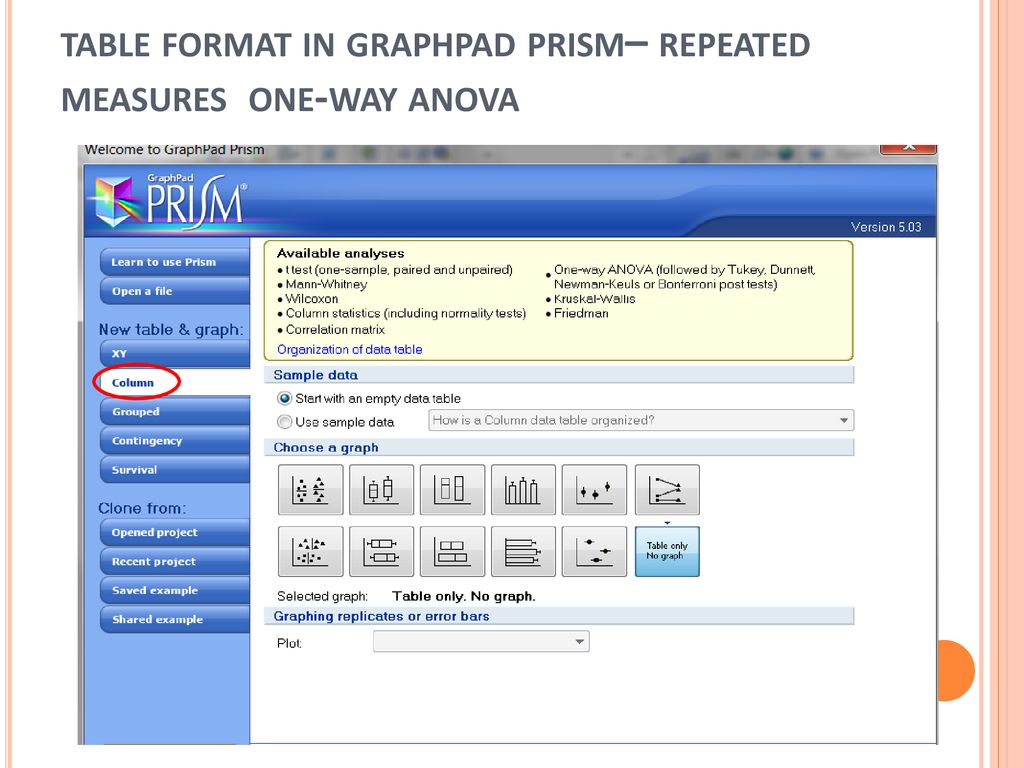
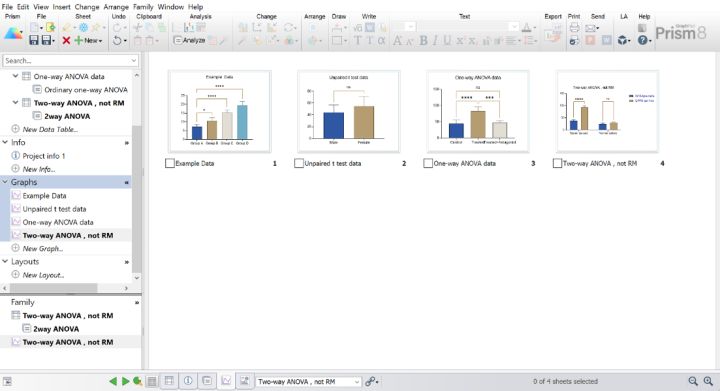
#Graphpad prism 6 教程 twoway anova how to
This chapter describes how to compute the Kruskal-Wallis test using the R software. It’s recommended when the assumptions of one-way ANOVA test are not met. Here we will compare the distribution of the evolutionary roots inferred for TFs and target genes using the Wilcoxon-Mann-Whitney test, and then generate violin plots (please refer to Trefflich et al. If the Kruskal–Wallis test is significant, a post-hoc analysis can be performed to determine which levels of the independent variable differ from each other level. When a significant site effect was found, a post hoc Mann–Whitney–Wilcoxon Rank Sum test using Bonferroni correction (from “Agricolae. In R, you would use wilcox.test(x, y, paired=FALSE) to run this test. Although there are many ANOVA experimental designs available, biologists are taught to pay special attention to the design of experiments, and generally make sure that the experiments are fully factorial (in the case of two-way or higher ANOVAs) and balanced.
